This is another welcome guest post from Gillian de Villiers, a Scientific Officer in our Vaccine Group. This was presented as a Journal Club article recently, and fit so well into my continuing theme of “viral diversity from water” that I asked her to write it up. Thanks Gillian!
Giant virus with a remarkable complement of genes infects marine zooplankton
Matthias G. Fischer, Michael J. Allen, William H. Wilson, and Curtis A. Suttle
PNAS published ahead of print October 25 2010 www.pnas.org/content/early/2010/10/15/1007615107
This publication covers the sequencing of the genome of Cafeteria roenbergensis virus(CroV). This nucleocytoplasmic large DNA virus (NCLDV) is the largest marine virus described to date, and its closest relative is Acanthamoeba polyphaga Mimivirus.
Among the questions raised in this paper are:
- what is the evolutionary origin of big viruses?
- Did they get their genes from horizontal gene transfer (including from eukaryotes), or
- are the “eukaryotic” genes viral in origin?
Spoiler alert: the authors do not answer this question.
Please note: this is a virus from a seawater host. It is the largest marine virus yet found, but how hard has anyone been looking? This ties in with Ed’s theme that we should be looking for viral diversity and interesting things in the water, because interesting things have been found there.
Some background…
This lytic virus strain was isolated off the coast of Texas in the 1990s. The host, Cafeteria roenbergensis was originally misidentified as a Bodo species. It is a major micro flagellate grazer (microzooplanton = major ocean predator) a 2-6um “bicoecid heterokant phagotrophic flagellate” and has been found in multiple marine environments including surface waters, deep sea sediment and hydrothermal vents.
In other words, the host is an extremely significant part of the ocean ecosystem, and has been found in most places. The authors note that protists host the largest viruses known and that other giant viruses probably are widespread in the oceans, but so far only the Acanthamoeba-infecting giant viruses have been characterised (Acanthamoeba does not live in the ocean). Viral infections of cyanobacteria play a significant role in global oxygen production; in a similar way the viral infections by CroV may have implications for carbon and other nutrient cycling and the “food chain” in the oceans, although this is beyond the scope of the article.]
Results
The genome is the second-largest viral genome described and at 730kb is very AT rich. Approximately 618kb is thought to be coding with 544 predicted protein-coding genes. At least 274 genes are expressed during infection. 22 percent of CroV CDSs (coding sequences) were probably best related to eukaryotic genes. Most CroV CDSs had unknown function, but 32% of CDSs could be assigned a putative function.
For enzymatic functions that have not previously been reported in any other viruses you can refer to Table S1 of the Supplemental materials.
This is similar to CroV’s closest known relative, Mimivirus, where of 911 predicted genes only 300 were assigned a predicted function (see table). Only 1/3 of their genes are common to these two viruses! This suggests tremendous diversity within the nucleocytoplasmic large DNA viruses, as they may have common evolutionary origins for some genes, but not for others. As viruses are not monophyletic (although the NCLDVs may be) and can be considered to be bags of protein that contain genetic material and share a strategy (rather than an origin) this may not be particularly surprising. But I find it amazing that so many potential genes, and so many unique potential genes, have been found in these organisms.
Included in the genes assigned function are genes involved in translation. CroV encodes an isoleucyl-tRNA synthetase and putative homologs of eukaryotic translation initiation factors. 22 tRNA genes and two putative tRNA-modifying enzymes: tRNA pseudouridine 5S synthase and tRNAIle lysidine synthetase were found. Mimivirus also has four tRNA synthetases and several putative translation factors.
| Cafeteria roenbergensis virus |
Acanthamoeba polyphaga Mimivirus |
| ~730kb dsDNA genome |
~1200kb dsDNA genome |
| 300nm capsid |
500-750nm capsid (publications differ) |
| Largest marine virus yet described |
Largest virus yet described |
| Second-largest virus yet described |
– |
| 544 predicted genes |
911 predicted genes |
| 174 genes with predicted function |
300 genes with predicted function |
| Host: Cafeteria roenbergensis |
Host: Acanthamoeba castellani (amoeba) |
| Habitat: marine environment |
Habitat: soil (?freshwater) |
| Genes shared with Mimivirus ~ 1/3 |
Genes shared with CroV ~ 1/5 |
Similarly to other large DNA viruses a number of DNA repair genes were found. This includes a base excision repair pathway that appears complete. In addition crov115’s gene product is predicted to be a CPD class 1 photolyase, the first viral homologue in its class. Crov149 appears to be part of a recently described photolyase/cytochrome group found in several bacterial phyla and euryarchaeotes, but not among established types of photolyase. The authors suggest that the only eukaryote with this gene, Paramecium tetraurelia may have acquired it by horizontal gene transfer from a giant virus
CroV also has transcription-related genes including eight DNA-dependent RNA polymerase II subunits, six transcription factors involved in transcription initiation, elongation, and termination, a tri-functional mRNA capping enzyme, a poly (A) polymerase, as well as helicases. Mimivirus provides considerably more genes for protein transcription and translation than most viruses, and sets up its own ‘virus factory’ in the cytoplasm of the cell. It is possible that CroV has a similar strategy, with viral gene transcription independent of the host and occurring in the cytoplasm.
Of the three DNA topoisomerases, two are very similar to the counterparts in Mimivirus. CroV TopoIB is the first viral homolog of the eukaryotic subfamily, but the Mimivirus TopoIB appears to be from the bacterial group. Although the evolutionary origin appears to differ, the topoisomerases are presumably important in transcription, translation or packaging of giant virus genomes, as they appear in both CroV and Mimivirus genomes.
CroV has four inteins: self-splicing proteins. They are found in DNA-dependent DNA polymerase B (PolB), TopoIIA, DNA-dependent RNA polymerase II subunit 2 (RPB2) and the large subunit of ribonucleotide reductase (RNR). Inteins have previously been found in viruses infecting eukaryotes, including Mimivirus PolB. CroV TopoIIA intein is the first case of an intein in a DNA topoisomerase gene.
Microarray analysis on the 12-18 hr infection cycle showed around half the predicted genes, and 63% of the tested genes were expressed during infection. Work on Mimivirus and PBSC-1 showed transcription of nearly all predicted genes, so this work may underestimate the true transcriptional activity of CroV. CroV gene expression has an early phase 0-3 hrs after infection affecting 150 genes, and a late phase affecting 124 genes 6 hrs or later post-infection including all the structural components predicted. A conserved early promoter motif “AAAAATTGA” was identified in 35% of CDSs and is nearly identical to the Mimivirus early promoter motif “AAAATTGA”. A promoter element for genes transcribed during the late phase of CroV infection was found that is unrelated to the putative late promoter motif in Mimivirus.
A genomic fragment involved in carbohydrate metabolism was also found. This 38kb fragment includes enzymes for biosynthesis of 3-deoxy-D-manno-octulosonate (KDO). This is part of the lipopolysaccharide layer in gram-negative bacteria and is found in the green alga Chlorella and the cell wall of higher plants. Ten of the enzymes involved in carbohydrate metabolism were expressed, suggesting a role in viral glycoprotein biosynthesis, suggesting the virion surface may be coated with KDO- or sialic acid-like glycoconjugates.
There are no homologs in Mimivirus suggesting this region must have been acquired after the CroV and Mimivirus lineages split (or that the Mimivirus lineage lost it subsequently?). This may have been acquired from bacteria, however GC content is even lower than for the rest of the CroV genome, and a number of the proteins are phylogenetically between bacterial and eukaryotic homologs.
Phylogenetics and Speculations
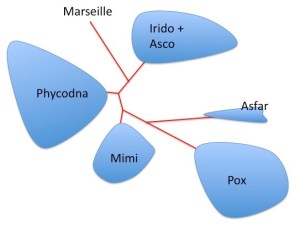
Phylogenetic reconstruction of NCLDV members. Redrawn and simplified from Fig. 4. The unrooted Bayesian Inference tree was generated from a 263-aa alignment of conserved regions of DNA polymerase B
CroV is an addition to the group of NCLDVs including Ascoviridae, Asfarviridae, Iridoviridae, Mimiviridae, Phycodnaviridae, Poxviridae and Marseillevirus, which are presumed to be monophyletic. CroV seems to be the closest known relative to Mimivirus although it is substantially smaller. The topology of the NCLDV tree strongly suggests the five largest viral genomes (all mimiviruses) are more closely related to each other than to other NCLDV families. They may have originated from an ancestral virus that was already an NCLDV that encoded more than 150 proteins.
Mimivirus is the most studied NCLDV, and is the largest. Most Mimivirus genes have no cellular homologs and may be very ancient, with 1/3 of genes having originated through gene and genome duplication and less than 15% of the genes having potentially been acquired by horizontal gene transfer from eukaryotes and bacteria. The CroV genome analysis is consistent with this view of giant virus evolution, with gene duplication and lineage-specific expansion contributing to the size of the CroV genome. The 38kb carbohydrate metabolism fragment may be a potential case of large-scale horizontal gene transfer from a bacterium. The PolB gene of CroV has high similarity with those of other marine isolates so it may represent a major group of marine viruses, that despite being virtually unknown have ecological significance.
CroV again shows overlap between large viruses and cellular life forms, adding to questions about the evolutionary history of giant viruses as well as what life itself is.