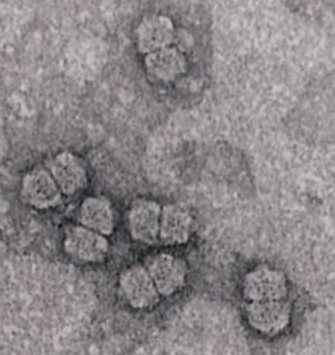
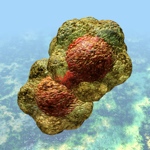
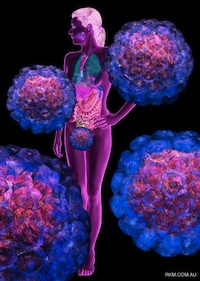
We thank Russell Kightley for permission to use the images
We thank Russell Kightley for permission to use the images
Marshall Bloom (Rocky Mountain Laboratories, NIAID) opened the plenary session on Thursday the 1st of December, with a talk on probing the pathogen-vector-host interface of tickborne flaviruses. Although thoroughly infected with a rhinovirus, he held our attention most ably while reminding us that while many flaviviruses are tick borne, the hard and soft body ticks that vector them are very phylogenetically different – as different as they are from spiders – meaning that if similar flaviruses replicated in them, these viruses may have much wider host range than we know.
He pointed out that while about 95% of the virus life cycle takes place in a tick, transmission to a vertebrate means suddenly adapting to a very different host. Infection in ticks is persistent, as befits their vector role – but vertebrate infection generally is not. It was interesting, as a sometime plant virologist, to hear that they look for dsRNA as a marker for replication, and do Ab staining for it: the technique was invented with plant viruses, and very few other virologists seem to appreciate that dsRNA can be quite easily isolated and detected.
They compared Vero and tick cells for virus replication, and saw significant differences: while tick cells could go out to 60+ days and look fine, Vero cells were severely affected at much shorter times post infection. There was also 100-fold less virus in tick cells, and prominent tubular structures in old infected tick cells. He noted that ticks evade host defences quite efficiently: eg they suppress host clotting during feeding, and there is huge gene activation in the tick during feeding. In another study to envy, they are doing array work on ticks to see what is regulated and how.
Linda Dixon (Institute for Animal Health, Pirbright, UK) recounted her lab’s work on African swine fever (ASFV), a poxvirus-like large DNA virus. The virus is endemic to much of Africa, and keeps escaping – and there is no effective vaccine to prevent spread, so regulation is by slaughter. There are 3 types of isolate, with the most highly pathogenic causing up to 10% fatality and a haemorrhagic syndrome. She described how in 2007 the virus had spread from Africa to Georgia, then in 2009 to southern Russia and all way to the far north, in wild boar.
There are more than 50 proteins in the dsDNA-containing virion; two infectious forms similar to the poxviruses with multilayer membranes and capsid layers can form, and neutralising Ab play no part in protection as a result. They studied the interaction of viruses with cells and the immune system, and compared the genomes of pathogenic and non-pathogenic strains, in order to understand how to develop an effective vaccine.
The biggest differences were large deletions in non-virulent isolates, including genes coding for proteins responsible for binding to RBC, and various immune evasion multicopy genes. They planned to target regions to delete to make an attenuated virus for vaccine. They had found non-essential genes involved in immune evasion, and ones that lower virulence, and had been systematically cutting them out. She noted that pigs can be protected if they survive natural infection and if vaccinated with TC-attenuated virus, and can be protected by passive transfer of Abs from immune pigs – which indicated that an effective live vaccine was very possible.
Subunit vaccines were being investigated, and they had found partial protection with baculovirus-expressed proteins. They were doing genome-wide screens for protective Ag, and were pooling Ags expressed from predicted ORFs in immunization trials – up to 47 Ags without reduction in specific T cell responses.

Discovery One
My former labmate Dion du Plessis (Onderstepoort Veterinary Institute, OVI) made a welcome return to Cape Town, with a talk entitled “2011: A Phage Odyssey”. He explained the title by noting the distinct resemblance of P1 coliphage to the Discovery One spacecraft dreamed up by Arthur C Clarke and Stanley Kubrick – and then went on to exuberantly and idiosyncratically recount a brief history of bacteriophages and their use in biotechnology since their discovery. A revelation from his talk was that the first discovery of phages was probably described by a gentleman named Hankin, in 1896 in Annales de l’Institut Pasteur: he

The 1896 paper from Annales de l'Institut Pasteur
showed that river water downstream of cholera-infested towns on the Jumma river in India contained no viable Cholera vibrio – and that this was a reliable property of the water. We were also introduced to the concept of turtles as undertakers in the Ganges….
He took us through the achievements of the Phage Group of Max Delbruck and others – where science was apparently fun, but also resulted in the establishment of modern molecular biology – through to the use of phages as exquisitely sensitive indicators immunochemistry studies in the 1960s.
All too soon we got to the modern uses of phages, with 3 types of gene library – random peptide, fragmented gene, and antibody V regions – being used to make recombinant phage tail proteins to be used for “panning” and enrichment purposes, in order to select either specific antibodies or antigens. Dion manages a research programme at OVI aimed at developing a new generation of veterinary vaccines – and has for some years now been making significant progress in generating reagents from a chicken IgY single-chain Fv phage display library.
Carolyn Williamson (IIDMM, UCT) gave us an update on CTL epitopes associated with control of HIV-1 subtype C infections. She said that it was now known that genome-wide association studies (GWAS) gives you certain HLAs which are associated with low viral load, and others with high – meaning that to some extent at least, control of infection was down to genetic luck. She noted that they and others had shown that CTL escape was quick: this generally happened in less than 5 weeks in acute phase infections.
They had looked for evidence of a fitness cost of CTL escape – and shown that it exists. She noted that this meant that even if one has “bad” HLA genes, if one was infected with a virus with fitness cost mutations from another, that one could still control infection.
It had been shown that “controllers” mainly have viruses with attenuating mutations, or have escapes in the p24 region – and it was a possible vaccine strategy to include these mutated epitopes in vaccines to help people with infections control their infections.
An interesting topic she broached was that of dual infections – there was the possibility of modelling if infection with two different viruses results in increased Ab neutralisation breadth, and if one would get different results if infections were staggered, possibly with increased nAb evolution if isolates were divergent. She noted it was possible to track recombination events with dual virus infections too.
It was interesting that, as far as Ab responses went, there were independent responses to 2 variants and one could get a boost in Ab titres to the superinfecting virus, but not a boost to Abs reacting with the originally-infecting virus
Carolyn was of the opinion that HIV vaccines needed to include CTL epitopes where escape is associated with fitness cost. She also reiterated that superinfection indicated that one can boost novel responses, which I take to mean that therapeutic applications are possible.
Ulrich Desselberger (University of Cambridge) is a long-time expert on rotaviruses and the vaccines against them, and it was a pleasure to finally hear him speak – and that he was mentoring young people in South Africa. He said that more than a third of children admitted to hospital worldwide were because of rotavirus infections, meaning that the viruses were still a major cause of death and morbidity – and they were ubiquitous.
He reviewed the molecular biology and replication cycle of rotaviruses in order to illustrate where they could be targeted for prevention of infection or therapy, and noted that drugs that interfere with lipid droplet homeostasis interfere with rotavirus replication because 2 viral proteins associated proteins of lipid droplets.
He stated that there were lots of recent whole-genome sequences – we already there were many types, based on the 2 virion surface proteins; we now know that other genes are also highly variable. As far as correlates of immunity were concerned, VP7 & 4 were responsible for eliciting neutralising Ab. Additionally, protective efficacy of VP6 due to elicitation of non-neutralising Ab had been shown in mice – but not in piglets, and not convincingly in humans. Abs to VP2, and NSP2 and 4 were also partially protective in humans. It was interesting that protection was not always correlated with high titre nAb responses.
He noted that in clinical disease primary infections partially protected against subsequent infections which are normally milder; subsequently no disease was seen even when infection occurred. Cross-protection occurred at least partially after initial infection, and this got better after more exposure. There was evidence one could get intracellular neutralisation by transcytosed Ab, and especially to VP6. Ab in the gut lumen was a good indication of protection.
As far as the live modern vaccines were concerned, Merck’s Rotateq elicited type-specific nAb, with 9% of recipients shedding live virus. GSK’s Rotarix gets elicits cross-reactive nAb and one gets 50% of recipients shedding virus.
While the vaccines seemed safe, he noted that where vaccines had been introduced, efficacy ranged from 90% in the USA and Europe, down to as low as 48% in Bangladesh, Malawi and SA, due to type mismatch, and that efficacy was correlated inversely with disease incidence and child mortality generally. He mentioned that there had been much VLP work, but that none of the candidates was near licensure.
Johan Burger (Stellenbosch University) spoke on one of the more important non-human virus problems in our immediate environment – specifically, those affecting wine grape production in our local area. He opened by stating that SA now produced 3.7% of the world’s wine, making grapes a nationally and especially locally important crop. Leafroll disease was a major worldwide problem – as well as being the reason for the wonderful autumn reddening seen in grapevines, it also significantly limited production in affected vineyards. His laboratory has done a lot of work in both characterising viruses in grapevine, and trying to engineer resistance to them. Lately they were also investigating the use of engineered miRNAs as a response to and means of controlling, virus infection.
His group has for a couple of years been involved in “metaviromic” or high-throughput sequencing studies of grapevines, with some significant success in revealing unsuspected infections. In this connection, he and Don Cowan pointed out that they had lots of data that they ignore – but which we should keep and study, as a resource for other studies not yet thought of.
As far as Johan’s work went, novel viruses kept popping up, including grapevine virus E (GVE), which hitherto had only been found in Japan. They were presently looking at Shiraz disease, which was unique to SA, and was still not understood. This was infectious, typified by a lack of lignification which led to rubbery vines, and kills plants in 5 years. It also limits the production of the eponymous grapes – a crime when SA shirazes seem to be doing so well!
Veterinary Virology and Vaccines parallel session.
I again dodged the clinical / HIV session because of my personal biases, and was again treated to a smorgasbord of delight: everyone spoke well, and to time, and I was really gratified to see so many keen, smart young folk coming through in South African virology. It was also very interesting to see highly topical subjects like Rift Valley fever and rare bunyavirus outbreaks being thoroughly covered, so I will concentrate on these.
P Jansen van Veeren (NICD, Johaanesburg) was again a speaker, this time representing his absent boss, Janusz Paweska. He gave an account of the 2010 Rift Valley fever outbreak in SA, and epidemiological findings in humans – something of keen interest to me. He said there had been some forecasting success for outbreaks in East Africa; however, there were long gaps between outbreaks, which were generally linked to abnormal rainfall and movement of mosquito and animal hosts. RVFV isolates differed in pathogenicity but were structurally and serologically indistinguishable – because virulence was due to the NSs protein, and not a virion component. He recounted how artificial flooding of a dambo in Kenya resulted in a population boom in the floodwater Aedes mosquitoes responsible for inititating an outbreak, and then of the Culex which maintained the epidemic. He said there was a strong correlation between viral load and disease severity.
In terms of South African epidemiology, there had been smaller outbreaks from 2008 round the Kruger National Park (NE SA), then in the Northern Cape and KZN in 2009. People had been infected from autopsy of animals, and handling butchered animal parts. The 2010 outbreak started in the central Free State after an unusually wet period, and had then spread to all provinces except Limpopo and KZN. In-house serological methods at the NICD were validated in-house too: these were HAI screening and IgM and IgG ELISAs and a virus neutralisation test. They had got 1600+ samples of human serum, and confirmed 242 cases of disease and 26 deaths for 2010.
He noted that with winter rains there was a continuous outbreak in the Western Cape, and in 2011 the epidemic had started again in the Eastern and Western Cape Provinces, but has since tailed off. Some 82% of human cases were people who occupationally handled dead animals, although there was some possibility of transmission by mosquitoes.
In human cases there was viraemia from 2-7 days, with IgM present transiently from 3 days at low level. They had sequenced partial GP2 after PCR from 47 isolates, and showed some recombination occurring. The 2010 isolates were very closely related to each other, and to a 2004 Namibian isolate. There had been no isolation from mosquitoes yet.
Two talks on FMDV followed: Belinda Blignaut (OVI and Univ Pretoria) spoke on indirect assessment of vaccine matching by serology, and Rahana Dwarka (OVI) on a FMDV outbreak in KZN Province in 2011. Belinda’s report detailed how 6 of 7 serotypes of FMDV occur in SA, with SAT-1 and -2 and O the most common – and that vaccines needed to be matched to emerging strains. This was done by indirect vaccine matching tests such as serological r-value, determined by the ratio of the reciprocal serum titre to the heterologous virus against that to the homologous virus. They had put 4 different viruses into cattle and got sera to test a range of 26 newly isolated viruses. While they had not got sequence from the test panel viruses, indications were that topotype 3 viruses are antigenically more disparate and that a vaccine consisting of topotype 1 or 2 antigens may not be effective in the control of FMD.
In introducing Rahana’s talk, the chair (Livio Heath, OVI) mentioned that there had been 5 different major animal pathogens causing outbreaks in SA over the last 3 years – and that they had to produce reagents and validate tests for ASFV, classical swine fever (CSF) and FMDV, etc, with each outbreak. Rahana described how they had neutralisation assays and blocking and competition ELISA for FMDV, as well as a big database of isolates from buffalo in KZN – so they were well-placed to type viruses found in cattle in the region.
C van Eeden (Univ Pretoria) had an intriguing account of their investigation of the occurrence of an orthobunyavirus causing neurological symptoms in horses and wildlife. Horses seem to be particularly vulnerable to many of the viruses involved in such disease, and so are a useful sentinel species. Shuni virus was first isolated from Culicoides midges and sheep and a child in Nigeria in the 1960s. SA workers subsequently found it in some livestock and Culex mosquitoes and in horses. The virus was shown to be a neurologic disease agent in horses and wildlife – then disappeared for some 30 years, much like Ebola. There is apparently a new research unit at UP with a BSL3 lab, so they are well equipped to do tests with the virus.
Ms van Eeden noted that the incidence of encephalitic disease in humans and animal in SA is underreported, and the causes are mainly unknown – a revelation to me! Horses are susceptible to many of the agents, and are useful sentinels – workers have identified flavi- and alphaviruses in some outbreaks, but many are not IDed. They had done cell culture and EM on samples from an ataxic horse: they got a bunyavirus-like virus by EM, and did bunya-specific PCR, and got Shuni virus back. Sequence relationships showed no linkage to type of animal or date, in subsequent samplings from horses, crocodiles, a rhino and a warthog, and from blood, brain and spinal cord. All positive wildlife were sampled in Limpopo Province; horses only from most other provinces.
She noted that latest cases were neurological, whereas previously these were mainly febrile. The virus accounted for 10% all neurological cases, with a 50% fatality rate. She noted further that vets often work without masks or gloves, and so had no protection from exposure in such cases…. There was no idea on what the vector was, but they would like to test mosquitoes, etc. Ulrich Desselberger suggested rodents may be a reservoir, but they don’t know if this is true.
Stephanie van Niekerk (Univ Pretoria) investigated alphaviruses as neurological disease agents in African wildlife. The most common alphaviruses in SA are Sindbis and Middelburg viruses. Old World alphaviruses are usually not too bad, and cause arthritic and febrile symptoms, while New World cause severe neurological diseases. Sindbis was been found in SA outbreaks in 1974. However, Stephanie noted that a severe neurological type had appeared since 2008 in horses. Accordingly, they looked at unexplained cases in wildlife in the period 2009-2011: brain and spinal cord samples were investigated for all cases. They found alphavirus in a number of rhinos, buffalo, warthog, crocodiles and jackal – and all except for one rhino were Middelburg virus. They want to isolate viruses in cell culture, and increase the size of regions used for cDNA PCR. Stephanie said the opinion was that the values of the animal involved justifies the development of vaccines.