I have often spoken of “molecular archeology” in my lectures, and of the possibility of identifying past epidemic / pandemic strains of human flu in particular, by looking at which viruses are recognised by antibodies from people who lived through the epidemics.
A new paper in Nature ups the stakes in this game considerably: a team led by one James E Crowe Jr describes how 32 survivors of the 1918 Spanish Flu pandemic – born in or before 1915 – were “mined” for antibodies, and seven donors additionally were shown to have circulating B cells which secreted antibodies which bound the 1918 H1N1 virus haemagglutinin (HA). The team isolated 5 monoclonal antibodies from these subjects, and showed that these potently neutralised the infectivity of the virus and bound the HA of a 1930 swine virus, but did not cross-react with the HAs of more recent human H1-containing viruses.

Neutralizing antibodies derived from the B cells of 1918 influenza pandemic survivors : Abstract : Nature via kwout
This achievement is undoubtedly a tour de force of modern molecular immunology – but is it useful?
Well, one very obvious fact is that people can obviously maintain significant levels of humoral immunity to viruses that infected them – in the words of the authors – “…well into the tenth decade of life.” This is good news indeed for vaccinees who received vaccines for viruses which do not change much, like measles, mumps and poliomyelitis viruses. However, given that influenza virus even of one H and N type can change so as to be unrecognisable in just a few years – the MAbs they generated did not react to any great extent with presumptively H1N1 human isolates from 1943, 1947, 1977 and 1999 – this is only of any use if the original virus were to be re-introduced somehow.
There was an intriguing statement in the paper which may shed some light on a long-running controversy as to the origin of the 1977 H1N1 pandemic, when the virus reappeared in humans for the first time since the early 1950s – allegedly as a result of an escape from a Soviet biowarfare lab.
“The 1F1 antibody bound and neutralized the 1977 virus, albeit to a lesser degree than either the 1918 or the Sw/30 viruses … and to a minimal degree the 1943 virus”.
Ye-e-e-sssss…strange, that. So the 1977 virus was antigenically more similar to 1930s era viruses than to one from 1943??
The proposed use of the findings also elicit biowar scenarios: for example, the fact that passive immunisation of people with antibodies to a particular virus can help them get over infection with it is purely academic for MAb to the 1918 virus – or is it?
I hope it is.
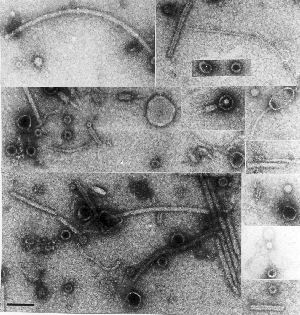
 About half the world’s oxygen is being produced by tiny photosynthesising creatures called phytoplankton in the major oceans. These organisms are also responsible for removing carbon dioxide from our atmosphere and locking it away in their bodies, which sink to the bottom of the ocean when they die, removing it forever and limiting global warming.
About half the world’s oxygen is being produced by tiny photosynthesising creatures called phytoplankton in the major oceans. These organisms are also responsible for removing carbon dioxide from our atmosphere and locking it away in their bodies, which sink to the bottom of the ocean when they die, removing it forever and limiting global warming. the viruses being very similar in genome organisation and indeed genome sequence.
the viruses being very similar in genome organisation and indeed genome sequence.