See on Scoop.it – Virology News
“To better understand the genetic diversity and genomic features of 41 coronaviruses (CoVs) identified from Kenya bats in 2006, seven CoVs as representatives of seven different phylogenetic groups identified from partial polymerase gene sequences, were subjected to extensive genomic sequencing. As a result, 15–16 kb nucleotide sequences encoding complete RNA dependent RNA polymerase, spike, envelope, membrane, and nucleocapsid proteins plus other open reading frames (ORFs) were generated. Sequences analysis confirmed that the CoVs from Kenya bats are divergent members of Alphacoronavirus and Betacoronavirus genera. Furthermore, the CoVs BtKY22, BtKY41, and BtKY43 in Alphacoronavirus genus and BtKY24 in Betacoronavirus genus are likely representatives of 4 novel CoV species. BtKY27 and BtKY33 are members of the established bat CoV species in Alphacoronavirus genus and BtKY06 is a member of the established bat CoV species in Betacoronavirus genus. The genome organization of these seven CoVs is similar to other known CoVs from the same groups except for differences in the number of putative ORFs following the N gene. The present results confirm a significant diversity of CoVs circulating in Kenya bats. These Kenya bat CoVs are phylogenetically distant from any previously described human and animal CoVs. However, because of the examples of host switching among CoVs after relatively minor sequence changes in S1 domain of spike protein, a further surveillance in animal reservoirs and understanding the interface between host susceptibility is critical for predicting and preventing the potential threat of bat CoVs to public health.”
We need to start worrying about MORE viruses from bats?? I knew those caves were a bad idea….
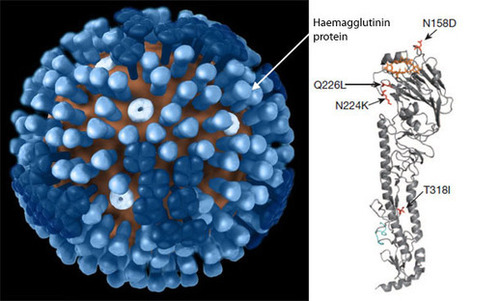
Graphic from Russell Kightley Media
See on www.sciencedirect.com