The news media are presently fascinated by the appearance of what looks like a new and nasty virus from my old home country, Zambia: see a link to The Times article of 7th October for the official word.
Which is “Don’t Panic”, written in large, friendly letters across the face of the newspaper….

From The Times article by Sashni Pather:
“Disease transmitted via bodily fluids
THE deputy director of the National Institute for Communicable Diseases has assured the public that there is no need to panic, despite the fact that four people have been killed by an unknown, highly contagious virus.
Doctor Lucille Blumberg, who also heads up the NICD’s epidemiology unit and consults to its special pathogens unit, referred to the death of Cecilia van Deventer, 36, as an “isolated case” and said test results were not yet available.
…A paramedic, Hannes Els, 33, who treated the critically ill Van Deventer in Zambia, and brought her to the Morningside Medi-Clinic in Sandton on September 12, died last Thursday after being infected with the highly contagious disease.
On Sunday a nurse and a cleaner, who had possibly been exposed to the disease, died.
Blumberg said: “The cause of death of the cleaner is still being investigated. We are busy conducting tests on all four deceased. The cleaner might not have been killed as a result of the virus.
“This is an isolated case. There have been no other reported cases in Lusaka, Zambia. “”
OK, so no obvious panic there, then – but disturbing echoes of another incident from 1996, when a Gabonese doctor was medevacced in to South Africa, and passed on an Ebola virus infection to a theatre nurse, Marilyn Lahana. He recovered, and she died – and there was close to panic in the land, as chronicled here. And here we are again…. Incidentally, anyone who wants to see how the first Ebola outbreaks of the electronic age unfolded can see a day-by-day history here, on my original Ebola pages. Still accessible, to my surprise!
That wonderful institution that is ProMED – who were the first people to break hard news of the Kikwit Ebola outbreak back in 1995, came to the attention of the serious medical reporting world as a result – has a slightly different view of the whole thing – and some interesting details not in the general news story. In this morning’s digest:
UNDIAGNOSED FATALITIES – SOUTH AFRICA ex ZAMBIA (02)
***********************************************
A ProMED-mail post
<http://www.promedmail.org>
ProMED-mail is a program of the International Society for Infectious Diseases
<http://www.isid.org>
[1]
Date: Mon 6 Oct 2008
Source: South African Broadcasting Corporation News online [edited]
<http://www.sabcnews.com/south_africa/health/0,2172,177844,00.htm>
A 4th person with viral haemorragic fever (VHF) symptoms has died. The virus has already claimed the lives of a Zambian national and 2 other people at the Morningside Clinic in Johannesburg. The woman was a cleaner at the clinic.
The National Health Department has issued an alert in Gauteng following these deaths. Unconfirmed tests indicate they may have died of [a] fatal viral haemorragic fever. An Outbreak Response and Tracking Team has been set up to contain any further spread. The department’s Zanele Mngadi says investigations are still underway into the cause of the deaths.
Mngadi confirmed the death of the 4th person, who was admitted at the Leratong hospital last night [5 Oct 2008]. The patient, who showed symptoms of VHF, was transferred to the Charlotte Maxeke Johannesburg
Academic Hospital, where she died. The health department says there is no need for South Africans to panic. The department’s Frew Denson says the fever is highly contagious but is only transmitted through body fluids.
It is reported that the virus can kill a person within 72 hours. VHF is an extremely infectious and life-threatening disease caused by [several different] viruses, including Ebola virus. The death rate [in the case of Ebola virus] can be as high as 90 percent. Symptoms vary but include fever, vomiting, diarrhea and bleeding.
Communicated by:
Rabelani Daswa <rabedaswa@gmail.com>
******
[2]
Date: Mon 6 Oct 2008
From: Amy Cantlay <inka@iwayafrica.com>
I have just read the posting (Undiagnosed fatalities – South Africa ex Zambia: RFI 20081005.3139) on your site, and it appears to be rather misleading. The chronological order of events (as I can gather) is as follows (None of this information has yet been confirmed.):
4 Sep 2008 – Index Case – female South African, (living in Zambia for many years) begins to suffer from flu-like symptoms.
9 Sep 2008 – She is slowly deteriorating. She sees multiple doctors in Lusaka.
11 Sep 2008 – She is admitted to hospital and deteriorates over night.
12 Sep 2008 – Paramedic is called in to evacuate her to South Africa. He does the transfer, along with another Dr assisting.
13 Sep 2008 – Index Case dies.
14 Sep 2008 Paramedic starts to develop flu-like symptoms.
14-27 Sep 2008 – Paramedic slowly deteriorates.
27 Sep 2008 – Paramedic is diagnosed as very sick and medivaced [sic] to South Africa. Nurse who treated Index case begins to get flu-like symptoms.
30 Sep 2008 – Paramedic dies.
1 Oct 2008 – Nurse who treated Index Case is admitted to hospital.
5 Oct 2008 – Nurse who treated Index Case dies.
The information that I can gather is the following:
1. Incubation period is as little as 2 days (paramedic), but as long as 14 days (nurse).
2. Disease course is generally 4-7 days of flu-like illness with patient only becoming critically ill in 2nd week of disease.
3. Further information is that Index Case reportedly had an eschar on one of her feet, thought to be from a tick-bite. She had also been in contact with horses from Congo in the weeks preceding her illness. Transmission is hypothesized to be by 2 means: tick-borne 1st (which may have brought the disease into the human population from the animal population) followed by direct contact with bodily fluids (resulting in human to human transmission).
4. It appears further hospital staff are now critically ill in Zambia, though this has not been confirmed.
5. If the incubation period is as long as 2 weeks, then we should still be closely watching all “contact-cases” for any signs of the disease. Those in contact with the Index case should be in the clear by now, while those in contact with the paramedic and the nurse (as well as any hospital staff who are currently sick) are still at high risk. One should probably work on a 21-day incubation period/quarantine period to be safe.
6. Chances are this is a new virus (or new subtype of virus) in the [family _Filoviridae_]. The only 2 known viruses in this group are Ebola and Marburg. It looks as though [the infection] may have entered Zambia from the Democratic Republic of the Congo (DRC) through a tick (carried on a horse), but again this cannot be confirmed.
This comment assumes that labs in South Africa have already tested all known VHFs. It is unlikely to be pneumonic plague, as this would have been discovered in South Africa; however, it is still a possibility that this [putative] viral disease has been in the Southern Province of Zambia (and that the 4 reported cases seen there were not diagnosed or wrongly called pneumonic plague).
7. The important steps in control are 1. effective quarantine of sick patients, and 2. monitoring of all “in-contact” cases, with quarantine as soon as any signs of flu or fever are noted. The government should also ideally make a statement to calm the panic and prevent people from fleeing the capital (potentially carrying the disease countrywide). This disease only spreads to people who are in very close contact with sick individuals. Those family members who are potentially incubating the disease should be encouraged to stay
around Lusaka, so that signs can be picked up quickly and treatment issued rapidly….[section on use of ribavirin – effective only against Lassa fever – edited out].
Communicated by:
Dr Amy Cantlay BVSc.MRCVS <inka@iwayafrica.com>
Veterinarian
Mkushi, Zambia
[ProMED-mail thanks Dr. Cantlay for her commentary, which contributes some interesting detail. At this point, it would not be useful to speculate further on the identity of the infectious agent responsible for the deaths of the 4 Zambian patients. No doubt a firm diagnosis will be available shortly from a South African reference laboratory. Several different viruses cause viral hemorrhagic fever. Of these, Ebola, Marburg, Lassa or Crimean-Congo hemorrhagic fever viruses have not been recorded in Zambia up to the present. A comprehensive account of these and other viruses responsible for hemorrhagic fevers can be found at the US CDC website: <http://www.cdc.gov/ncidod/diseases/virlfvr/virlfvr.htm>.
So: an unknown fever-causing agent, possibly associated with a tick bite in the index case, but which seems definitely to be transmitted quite efficiently via exposure to (presumably) body fluids, in a hospital setting…which does not appear to be known strains / types of Marburg or Ebola viruses, or Crimean-Congo haemorrhagic fever virus.
Loose in Johannesburg…part of a greater conurbation housing some 9 million people….
Should we be worried??
And the answer would be – NO.
If Ebola didn’t spread out of Kikwit in 1995 – a city of 500 000+ with no decent infrastructure to speak of – even to get as far as Kinshasa, then why should it spread in Johannesburg, or even Lusaka, where the infrastructure is MUCH more sophisticated?
I will leave this with a couple of quotes from posts I compiled on Ebola back in August 1995 from the fondly-remembered virology group at bio.net:
“To: virology@net.bio.net
From: ED@molbiol.uct.ac.za (“Ed Rybicki”)
Subject: Re: The Ebola virus – the end of the civilized world
Date: 18 Aug 1995 05:53:07 -0700
I would say you – and many others – are being unnecessarily
frightened by a concerted media campaign designed at selling lurid
books and films. Listen – for a change – to what experts tell you,
and react accordingly.
That is, RELAX!!!!!”
And:
“To: virology@net.bio.net
From: york@mbcrr.harvard.edu (Ian A. York)
Subject: Re: Ebola: the greatest threat, continued
In article <99792FA2E69@ida.ruc.dk>, wrote:
>
>Ebola is another ballgame. There is no way to protect effectively
>against this disease, and we have seen at mutation of this virus, Ebola
>Reston, that evidently was airborn (luckily it only affects monkeys).
…
Ebola has killed less than 400 people in the past decade. By contrast, typhoid fever kills over 600,000 people per year; measles kills 1,000,000 (one million) people per year. If you think Ebola has the potential tokill anywhere near that many, you don’t understand the virus. The Ebola outbreak in Kikwit *was* the worst-case scenario; *everything* went wrong. 300 deaths. Not trivial. But a tiny fraction of the real killers.
Lobby and try to get measles vaccine in Africa, if you want to do some good. So don’t waste your time worrying about Ebola.”
Amen to that! Pity we have to keep revisiting The Threat From Darkest Africa – maybe we can sell the rest of the world some vaccines against them sometime soon…B-)






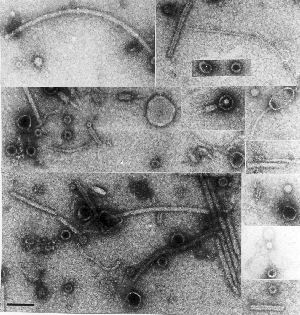
 West Nile virus
West Nile virus 
