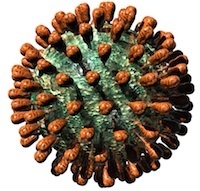
The following text has now appeared in modified form in an ebook, for sale for US$4.99 on the iBooks Store
The Ultracentrifuge, Eggs and Flu
The ultracentrifuge
A technical development that was to greatly advance the study of viruses was begun in 1923, but only reached fruition by the 1930s: this was the ultracentrifuge, invented and developed first by Theodor (“The”) Svedberg in Sweden as a purely analytical tool, and later by JW Beams and EG Pickels in the USA as an analytical and preparative tool. The ultracentrifuge revolutionised first, the physical analysis of proteins in solution, and second, the purification of proteins, viruses and cell components, by allowing centrifugation at speeds high enough to allow pelleting of subcellular fractions.
Analytical centrifugation and calculation of molecular weights of particles gave some of the first firm evidence that certain proteins, and virus particles, were large, regular objects. Indeed, it came to be taken as a given that one of the fundamental properties of a virus particle was its sedimentation coefficient, measured in svedbergs (a unit of 10-13 seconds, shown as S20,W). This is also how ribosomes of pro- and eukaryotes came to be named: these are known as 70S (prokaryote) and 80S ribosomes, respectively, based on their different sedimentation rates.
The Official Discovery of Influenza Virus
In 1931, Robert Shope in the USA managed to recreate swine influenza by intranasal administration of filtered secretions from infected pigs. Moreover, he showed that the classic severe disease required co-inoculation with a bacterium – Haemophilus influenza suis – originally thought to be the only agent. He also pointed out the similarities between the swine disease and the Spanish Flu, where most patients died of secondary infections. However, he also suggested that the virus survived seasonally in a cycle involving the pig, lungworms, and the earthworm, which is now known to be completely wrong.
This notwithstanding, he found that people who had survived infection during the 1918 pandemic had antibodies protecting them against the swine flu virus, while people born after 1920 did not, which showed that the 1918 human and swine flu viruses were very similar if not identical. This was a very relevant discovery for what happened much later, in the 2009 influenza pandemic, when the same virus apparently came back into the human population from pigs after circulating in them continuously since 1918.
Shope went on in 1932 to discover, with Peyton Rous, what was first called the Shope papillomavirus and later Cottontail rabbit papillomavirus: this causes benign cancers in the form of long hornlike growths on the head and face of the animal. This may explain the sightings in the US Southwest of the near-mythical “jackalope”.

Influenza viruses in pigs
Patrick Laidlaw and William Dunkin, working in the UK at the National Institute for Medical Research (NIMR), had by 1929 successfully characterised the agent of canine distemper – a relative of measles, mumps and distemper morbilliviruses – as a virus, proved it infected dogs and ferrets, and in 1931 got a vaccine into production that protected dogs. This was made from chemically inactivated filtered tissue extract from infected animals. Their work built on and completely eclipsed earlier findings, such as those of Henri Carré in France in 1905, who first claimed to have shown it was a filterable agent, and Vittorio Puntoni, who first made a vaccine in Italy from virus-infected brain tissue inactivated with formalin in 1923.
Influenza and Ferrets: the Early Days
Continuing from Laidlaw and Dunkin’s work in the same institute, Christopher Andrewes, Laidlaw and W Smith reported in 1933 that they had isolated a virus from humans infected with influenza from an epidemic then raging. They had done this by infecting ferrets with filtered extracts from infected humans – after the fortuitous observation that ferrets could apparently catch influenza from infected investigators! The “ferret model” was very valuable – see here for modern use of ferrets – as strains of influenza virus could be clinically distinguished from one another.
Eggs and Flu and Yellow Fever

Influenza virus and eggs: large-scale culture
Frank Macfarlane Burnet from Australia visited the NIMR in the early 1930s, and learned a number of techniques he used to great effect later on. Principal among these was the technique of embryonated egg culture of viruses – which he took back to Melbourne, and applied to the infectious laryngotracheitis virus of chickens in 1936. This is a herpesvirus, first cultivated by JR Beach in the USA in 1932: Burnet used it to demonstrate that it was possible to do “pock assays” on chorioallantoic membranes that were very similar to the plaque assays done for bacteriophages, with which he was also very familiar. Also in 1936, Burnet started a series of experiments on culturing human influenza virus in eggs: he quickly showed that it was possible to do pock assays for influenza virus, and that
“It can probably be claimed that, excluding the bacteriophages, egg passage influenza virus can be titrated with greater accuracy than any other virus.”
Max Theiler and colleagues in the USA took advantage of the new method of egg culture to adapt the French strain of yellow fever virus (YFV) he had grown in mouse brains to being grown in chick embryos, and showed that he could attenuate the already weakened strain even further – but it remained “neurovirulent”, as it caused encephalitis or brain inflammation in monkeys. He then adapted the first YFV characterised – the Asibi strain, from Ghana in 1927 – to being grown in minced chicken embryos lacking a spinal cord and brain, and showed in 1937 that after more than 89 passages, the virus was no longer “neurotrophic”, and did not cause encephalitis. The new 17D strain of YFV was successfully tested in clinical trials in Brazil in 1938 under the auspices of the Rockefeller Foundation, which has supported YFV work since the 1920s. The strain remains in use today, and is still made in eggs.
Virus purification and the physicochemical era
Given that the nature of viruses had prompted people to think of them as “chemical matter”, researchers had attempted from early days to isolate, purify and characterise the infectious agents. An early achievement was the purification of a poxvirus in 1922 by FO MacCallum and EH Oppenheimer.
Much early work was done with bacteriophages and plant viruses, as these were far easier to purify or extract at the concentrations required for analysis, than animal or especially human viruses.
CG Vinson and AM Petre, working with the infectious agent causing mosaic disease in tobacco – tobacco mosaic virus, or TMV – showed in 1931 that they could precipitate the virus from suspension as if it were an enzyme, and that infectivity of the precipitated preparation was preserved. Indeed, in their words:
“…it is probable that the virus which we have investigated reacted as a chemical substance”.
Viruses in Crystal
An important set of discoveries started in 1935, when Wendell Stanley in the USA published the first proof that TMV could be crystallised, at the time the most stringent way of purifying molecules. He also reported that the “protein crystals” were contaminated with small amounts of phosphorus. An important finding too, using physical techniques including ultracentrifugation and later, electron microscopy, was that the TMV “protein” had a very high molecular weight, and was in fact composed of large, regular particles. This was a very significant discovery, as it indicated that some viruses at least really were very simple infectious agents indeed.

TMV particle: 95% protein, 5% RNA
However, his conclusion that TMV was composed only of protein was soon challenged, when Norman Pirie and Frederick Bawden working in the UK showed in 1937 that ribonucleic acid (RNA) – which consists of ribose sugar molecules linked by phosphate groups – could be isolated consistently from crystallised TMV as well as from a number of other plant viruses, which accounted for the phosphorus “contamination”. This resulted in the realisation that TMV and other plant virus particles – now known to be virions – were in fact nucleoproteins, or protein associated with nucleic acid.
Stanley received a share of the Nobel Prize in Chemistry in 1946 for his work on TMV: it is instructive to read his acceptance speech from the time to realise what the state of the science that was becoming virology was at the time. He wrote:
“Since the original discovery of this infectious, disease-producing agent, known as tobacco mosaic virus, well over three hundred different viruses capable of causing disease in man, animals and plants have been discovered. Among the virus-induced diseases of man are smallpox, yellow fever, dengue fever, poliomyelitis, certain types of encephalitis, measles, mumps, influenza, virus pneumonia and the common cold. Virus diseases of animals include hog cholera, cattle plague, foot-and-mouth disease of cattle, swamp fever of horses, equine encephalitis, rabies, fowl pox, Newcastle disease of chickens, fowl paralysis, and certain benign as well as malignant tumors of rabbits and mice. Plant virus diseases include tobacco mosaic, peach yellows, aster yellows, potato yellow dwarf, alfalfa mosaic, curly top of sugar beets, tomato spotted wilt, tomato bushy stunt, corn mosaic, cucumber mosaic, and sugar cane yellow stripe. Bacteriophages, which are agents capable of causing the lysis of bacteria, are now regarded as viruses”.
Two of the most interesting things about the article, however, are the electron micrographs of virus particles – Stanley had one of the first electron micrsoscopes available at the time – and the table of sizes of viruses, proteins and cells that had been determined by then by techniques such as ultracentrifugation and filtration: TMV was known to be rodlike, 15 x 280 nm; vaccinia was 210 x 260 nm; poliomyelitis was 25 nm; phages like T2 were known to have a head-and-tail structure.
Seeing is Believing: the Electron Microscope
The development of the electron microscope, in Germany in the 1930s, represented a revolution in the investigation of virus structures: while virions of viruses like variola and vaccinia could just about be seen by light microscopy – and had been, as early as 1887 by John Buist and others – most viruses were far too small to be visualised in this way.
While Ernst Ruska received a Nobel Prize in 1986 for developing the electron microscope, it was his brother Helmut who first imaged virus particles – using beams of electrons deflected off virus particles coated in heavy metal atoms. From 1938 through the early 1940s, using his “supermicroscope”, he imaged virions of poxviruses, TMV, varicella-zoster herpesvirus, and bacteriophages, and showed that they were all particulate – that is, they consisted of regular and sometimes complex particles, and were often very different from one another. He even proposed in 1943 a system of viral classification on the basis of their perceived structure.
While electron microscopy was also used medically to some extent thereafter – for example, in differentiating smallpox from chickenpox by imaging particles of variola virus and varicella-zoster virus, respectively, derived from patients’ vesicles – its use was limited by the expense and cumbersome nature of sample preparation. For example, the micrographs in Stanley’s 1946 paper were all done with samples “…prepared with gold by the shadow-casting technique”.
The use of the cumbersome technique of metal shadow-casting, and the highly inconvenient nature of electron microscopy as a routine tool all changed from 1959 onwards, when Sydney Brenner and Robert Horne published “A negative staining method for high resolution electron microscopy of viruses”. This method involves the use of viruses in liquid samples deposited on carbon-coated metal grids, and then stained with heavy-metal salts such as phosphotungstic acid (PTA) or uranyl acetate.
This simple technique revolutionised the field of electron microscopy, and within just a few years much information was acquired about the architecture of virus particles. Not only were the overall shapes of particles revealed, but also the details of the symmetrical arrangement of their components. Some beautiful examples can be seen here, at the Cold Spring Harbor site.

Depiction of the effects of using a heavy metal salt solution to negatively stain particles on a carbon film. The stain (dark) pools around the particles (light). Human rotavirus particles, stained from below (left) and by immersion (right).
Images copyright LM Stannard
Click here for Part 1: Filters and Discovery
here for Part 3: Phages, Cell Culture and Polio
and here for Part 4: RNA Genomes and Modern Virology
Copyright Edward P Rybicki and Russell Kightley, February 2015, except where otherwise noted.